随着PTE考生对PTE口语和PTE听力的重视,大家口语和听力的分数得到极大提高,但是PTE阅读渐渐成为考生们新的难题。
墨尔本悉尼文波PTE特别为PTE考生们挑选了适合练习PTE阅读的文章,主题,内容,长度都与PTE阅读题中的文章相似。激活学过的词汇,更新新的词汇,提高阅读速度,全面提升自己的阅读能力。
ABOUT 120,000 types of protein molecule have yielded up their structures to science. That sounds a lot, but it isn’t. The techniques, such as X-ray crystallography and nuclear-magnetic resonance (NMR), which are used to elucidate such structures do not work on all proteins. Some types are hard to produce or purify in the volumes required. Others do not seem to crystallise at all—a prerequisite for probing them with X-rays. As a consequence, those structures that have been determined include representatives of less than a third of the 16,000 known protein families. Researchers can build reasonable computer models for around another third, because the structures of these resemble ones already known. For the remainder, however, there is nothing to go on.
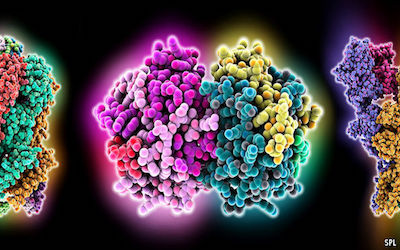
In addition to this lack of information about protein families, there is a lack of information about those from the species of most interest to researchers: Homo sapiens. Only a quarter of known protein structures are human. A majority of the rest come from bacteria. This paucity is a problem, for in proteins form and function are intimately related. A protein is a chain of smaller molecules, called amino acids, that is often hundreds or thousands of links long. By a process not well understood, this chain folds up, after it has been made, into a specific and complex three-dimensional shape. That shape determines what the protein does: acting as a channel, say, to admit a chemical into a cell; or as an enzyme to accelerate a chemical reaction; or as a receptor, to receive chemical signals and pass them on to a cell’s molecular machinery. (Models of all three, in that order, are shown above.)
crystallography /ˌkrɪstə’lɒgrəfɪ/ 晶体学
resonance /’rez(ə)nəns/ n. 回响,回荡
elucidate /ɪ’l(j)uːsɪdeɪt/ v. 阐明
paucity /’pɔːsɪtɪ/ n. 不足